Multi-angle fusion on reference image (non-iterative)¶
In this notebook, we explain and illustrate how-to performs multi-angle registration and fusion.
%matplotlib inline
import matplotlib.pyplot as plt
from os.path import join
import numpy as np
from timagetk.algorithms.blockmatching import blockmatching
from timagetk.algorithms.pointmatching import pointmatching
from timagetk.algorithms.trsf import apply_trsf
from timagetk.algorithms.trsf import compose_trsf
from timagetk.algorithms.trsf import inv_trsf
from timagetk.algorithms.trsf import trsfs_averaging
from timagetk.algorithms.averaging import images_averaging
from timagetk.algorithms.resample import resample
from timagetk.algorithms.trsf import create_trsf
from timagetk.components.spatial_image import SpatialImage
from timagetk.io import imread
from timagetk.io import imsave
from timagetk.io.image import _image_from_url
from timagetk.components.multi_channel import combine_channels
from timagetk.visu.mplt import grayscale_imshow
/builds/J-gEBwyb/0/mosaic/timagetk/src/timagetk/components/labelled_image.py:31: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from tqdm.autonotebook import tqdm
def imshow(ax, spim, title=""):
spim_extent = [-0.5, spim.get_extent('x') - 0.5, spim.get_extent('y') - 0.5, -0.5]
fig = ax.imshow(spim, cmap='gray', extent=spim_extent, vmin=0, vmax=255)
ax.xaxis.tick_top()
if title != "":
ax.set_title(title)
else:
try:
ax.set_title(f"{spim.filename}")
except:
pass
return fig
def add_landmarks(ax, ldmks):
px, py, c = np.array([(p[0], p[1], n) for n, p in enumerate(ldmks)]).T
ax.scatter(px, py, c=c, cmap='Set3', ec='k', s=80)
Load the multi-angle images:¶
img_list = [imread(fname) for fname in fnames]
Down-sample the images to speed-up the computations¶
print("# - Voxelsize (ZYX) for the initial image list:")
print("\n".join([f"Image '{im.filename}' (#{n}): {im.voxelsize}" for n, im in enumerate(img_list)]))
print("# - Shape (ZYX) for the initial image list:")
print("\n".join([f"Image '{im.filename}' (#{n}): {im.shape}" for n, im in enumerate(img_list)]))
# - Voxelsize (ZYX) for the initial image list:
Image '090223-p58-flo-top.lsm' (#0): [1.0, 0.20031953747828882, 0.20031953747828882]
Image '090223-p58-flo-tilt1.lsm' (#1): [1.0, 0.20031953747828882, 0.20031953747828882]
Image '090223-p58-flo-tilt2.lsm' (#2): [1.0, 0.20031953747828882, 0.20031953747828882]
# - Shape (ZYX) for the initial image list:
Image '090223-p58-flo-top.lsm' (#0): (59, 460, 460)
Image '090223-p58-flo-tilt1.lsm' (#1): (50, 460, 460)
Image '090223-p58-flo-tilt2.lsm' (#2): (54, 460, 460)
img_list = [resample(im, voxelsize=[1., 0.5, 0.5]) for im in img_list]
INFO [timagetk.algorithms.resample] Resampling using provided voxel-sizes: [1.0, 0.5, 0.5]
INFO [timagetk.algorithms.resample] Resampling using provided voxel-sizes: [1.0, 0.5, 0.5]
INFO [timagetk.algorithms.resample] Resampling using provided voxel-sizes: [1.0, 0.5, 0.5]
print("# - Voxelsize (ZYX) for the resampled image list:")
print("\n".join([f"Image '{im.filename}' (#{n}): {im.voxelsize}" for n, im in enumerate(img_list)]))
print("# - Shape (ZYX) for the resampled image list:")
print("\n".join([f"Image '{im.filename}' (#{n}): {im.shape}" for n, im in enumerate(img_list)]))
# - Voxelsize (ZYX) for the resampled image list:
Image '090223-p58-flo-top.lsm' (#0): [1.0, 0.5, 0.5]
Image '090223-p58-flo-tilt1.lsm' (#1): [1.0, 0.5, 0.5]
Image '090223-p58-flo-tilt2.lsm' (#2): [1.0, 0.5, 0.5]
# - Shape (ZYX) for the resampled image list:
Image '090223-p58-flo-top.lsm' (#0): (59, 184, 184)
Image '090223-p58-flo-tilt1.lsm' (#1): (50, 184, 184)
Image '090223-p58-flo-tilt2.lsm' (#2): (54, 184, 184)
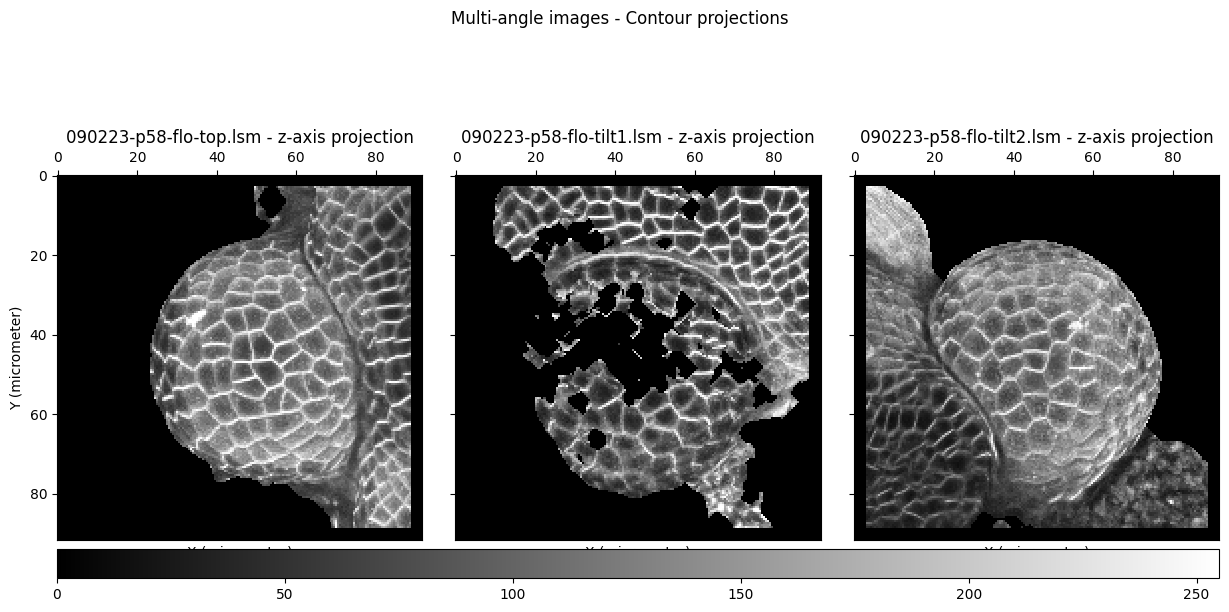
Intensity projections for multi-angle images¶
f = grayscale_imshow(img_list, suptitle="Multi-angle images - Contour projections", title=[img.filename for img in img_list], val_range=[0, 255], threshold=40)
0%| | 0/3 [00:00<?, ?image/s]
33%|███▎ | 1/3 [00:03<00:07, 3.58s/image]
WARNING [guess_intensity_threshold] Could not detect intensity threshold with 'bimodal search'...
INFO [guess_intensity_threshold] Using value obtained from 'percentile search'...
67%|██████▋ | 2/3 [00:07<00:03, 3.84s/image]
100%|██████████| 3/3 [00:11<00:00, 3.94s/image]
100%|██████████| 3/3 [00:11<00:00, 3.89s/image]
Initialisation using manually defined landmarks¶
Defines the reference image¶
We use the first image of the list, which is a ‘top’ view, as the initial reference image.
ref_img = img_list[0]
Load the manual registration landmarks:¶
As the angles between the “views” is important, we start with a manual initialisation using landmarks.
flo_pts_01 = np.loadtxt('p58_t0_floating_ldmk-01.txt')
flo_pts_02 = np.loadtxt('p58_t0_floating_ldmk-02.txt')
ref_pts_01 = np.loadtxt('p58_t0_reference_ldmk-01.txt')
ref_pts_02 = np.loadtxt('p58_t0_reference_ldmk-02.txt')
Creates manual initial transformations with pointmatching algorithm:¶
man_trsf_01 = pointmatching(flo_pts_01, ref_pts_01, template_img=ref_img, method='rigid')
man_trsf_02 = pointmatching(flo_pts_02, ref_pts_02, template_img=ref_img, method='rigid')
init_trsfs = [man_trsf_01, man_trsf_02]
Visualisation of the manual rigid transformations¶
To assess the quality of the manual rigid transformations we apply the previously computed transformations obtained from pointmatching to the floating images.
man_reg_imgs = [ref_img]
for i, man_trsf in enumerate(init_trsfs):
man_reg_imgs.append(apply_trsf(img_list[i+1], man_trsf, template_img=ref_img))
Then, we represent one slice of the intensity signal of each image in a different color to appreciate the “signal alignment” (or not).
The corresponding color code:
090223-p58-flo-top.lsm: red090223-p58-flo-tilt1.lsm: green090223-p58-flo-tilt2.lsm: blue
z_slices = [5, 10, 15, 20]
fig, axes = plt.subplots(1, 4)
fig.set_size_inches(15, 5)
fig.set_dpi(160)
fig.suptitle("Manual rigid multi-angle registration")
blend = combine_channels(man_reg_imgs, colors=['red', 'green', 'blue'])
for n, zsl in enumerate(z_slices):
imshow(axes[n], blend[zsl, :, :], title = f"z-slice #{zsl}")
fig.subplots_adjust(wspace=0, hspace=0)
fig.tight_layout()
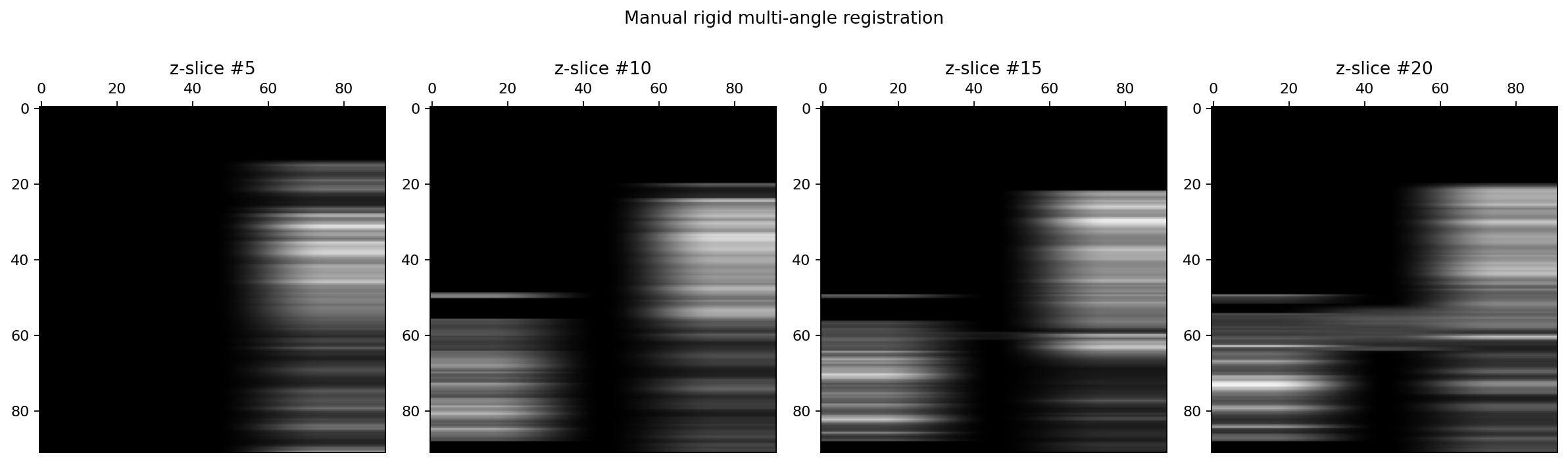
As we can see, the manual registration could be better as the green (090223-p58-flo-tilt1.lsm) and blue (090223-p58-flo-tilt2.lsm) channels do not correctly align with the reference (red) channel (090223-p58-flo-top.lsm).
To improve this initial registration we performs a rigid registration of the two floating images onto the reference using these manual tranformations.
Improve manual initial transformations with blockmatching algorithm:¶
for flo_idx, flo_img in enumerate(img_list[1:]):
man_trsf = blockmatching(flo_img, ref_img, method='rigid',
init_trsf=init_trsfs[flo_idx], pyramid_lowest_level=2)
init_trsfs[flo_idx] = man_trsf
INFO [timagetk.algorithms.blockmatching] RIGID registration with an initialisation matrix
INFO [timagetk.algorithms.blockmatching] RIGID registration with an initialisation matrix
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
Visualisation of the initial rigid transformations¶
man_reg_imgs = [ref_img]
for i, init_trsf in enumerate(init_trsfs):
man_reg_imgs.append(apply_trsf(img_list[i+1], init_trsf, template_img=ref_img))
Then, we represent one slice of the intensity signal of each image in a different color to appreciate the “signal alignment” (or not).
The corresponding color code:
090223-p58-flo-top.lsm: red090223-p58-flo-tilt1.lsm: green090223-p58-flo-tilt2.lsm: blue
z_slices = [5, 10, 15, 20]
fig, axes = plt.subplots(1, 4)
fig.set_size_inches(15, 5)
fig.set_dpi(160)
fig.suptitle("Initial rigid multi-angle registration")
blend = combine_channels(man_reg_imgs, colors=['red', 'green', 'blue'])
for n, zsl in enumerate(z_slices):
imshow(axes[n], blend[zsl, :, :], title = f"z-slice #{zsl}")
fig.subplots_adjust(wspace=0, hspace=0)
fig.tight_layout()
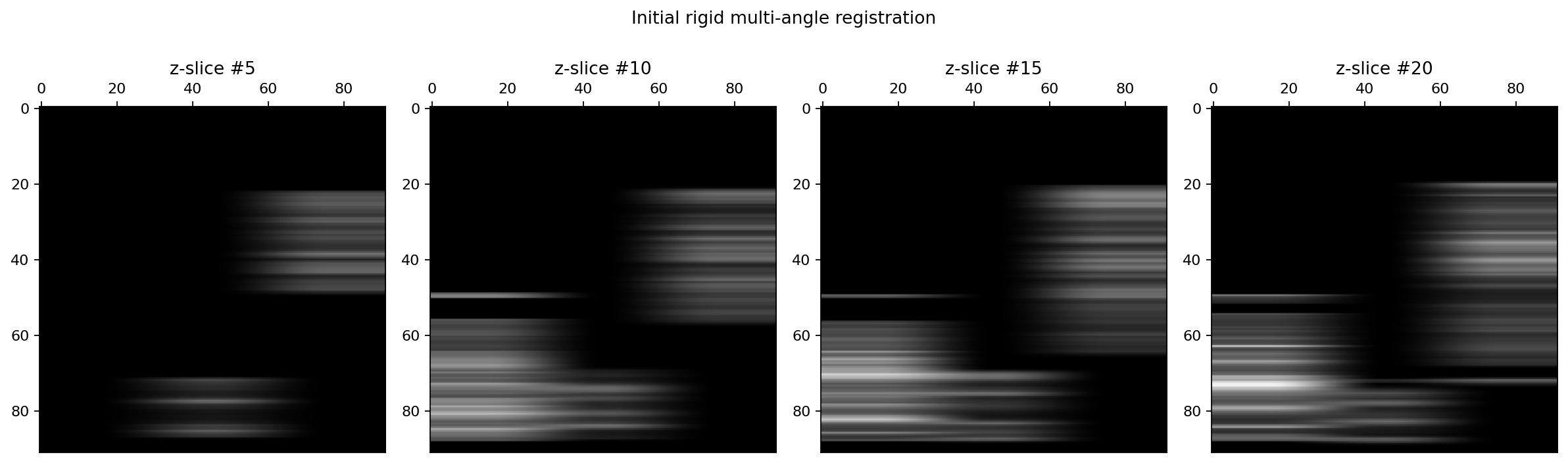

As we can see on the previous figure, the signal are now much better aligned and we may now proceed to the fusion of the two floating images onto the reference image.
Fusion on reference image¶
To improve the quality of the signal registration prior to the image fusion step, we compute the affine and non-linear transformations of the floating images onto the reference image.
Affine & non-linear deformations estimations¶
lin_trsfs = [] # linear part of the transformations
vf_trsfs = [] # non-linear part of the transformation
for flo_idx, flo_img in enumerate(img_list[1:]):
# Estimate the AFFINE transformation from floating image to reference image using manual initialisation:
lin_trsf = blockmatching(flo_img, ref_img, method='affine', left_trsf=init_trsfs[flo_idx], pyramid_lowest_level=1)
lin_trsfs.append(lin_trsf)
# Estimate the VECTORFIELD transformation from floating image to reference image:
vf_trsf = blockmatching(flo_img, ref_img, method='vectorfield', left_trsf=compose_trsf([init_trsfs[flo_idx], lin_trsf], template_img=ref_img), pyramid_lowest_level=1)
vf_trsfs.append(vf_trsf)
INFO [timagetk.algorithms.blockmatching] AFFINE registration with a 'left' initialisation matrix
INFO [timagetk.algorithms.blockmatching] VECTORFIELD registration with a 'left' initialisation matrix
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
- processing level # 1
- Iteration # 1 Level # 1 Size 128x 128x 1
- Iteration # 2 Level # 1 Size 128x 128x 1
- Iteration # 3 Level # 1 Size 128x 128x 1
- Iteration # 4 Level # 1 Size 128x 128x 1
- Iteration # 5 Level # 1 Size 128x 128x 1
- Iteration # 6 Level # 1 Size 128x 128x 1
- Iteration # 7 Level # 1 Size 128x 128x 1
- Iteration # 8 Level # 1 Size 128x 128x 1
- Iteration # 9 Level # 1 Size 128x 128x 1
- Iteration # 10 Level # 1 Size 128x 128x 1
INFO [timagetk.algorithms.blockmatching] AFFINE registration with a 'left' initialisation matrix
INFO [timagetk.algorithms.blockmatching] VECTORFIELD registration with a 'left' initialisation matrix
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
- processing level # 1
- Iteration # 1 Level # 1 Size 128x 128x 1
- Iteration # 2 Level # 1 Size 128x 128x 1
- Iteration # 3 Level # 1 Size 128x 128x 1
- Iteration # 4 Level # 1 Size 128x 128x 1
- Iteration # 5 Level # 1 Size 128x 128x 1
- Iteration # 6 Level # 1 Size 128x 128x 1
- Iteration # 7 Level # 1 Size 128x 128x 1
- Iteration # 8 Level # 1 Size 128x 128x 1
- Iteration # 9 Level # 1 Size 128x 128x 1
- Iteration # 10 Level # 1 Size 128x 128x 1
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
- processing level # 1
- Iteration # 1 Level # 1 Size 128x 128x 1
- Iteration # 2 Level # 1 Size 128x 128x 1
- Iteration # 3 Level # 1 Size 128x 128x 1
- Iteration # 4 Level # 1 Size 128x 128x 1
- Iteration # 5 Level # 1 Size 128x 128x 1
- Iteration # 6 Level # 1 Size 128x 128x 1
- Iteration # 7 Level # 1 Size 128x 128x 1
- Iteration # 8 Level # 1 Size 128x 128x 1
- Iteration # 9 Level # 1 Size 128x 128x 1
- Iteration # 10 Level # 1 Size 128x 128x 1
- processing level # 3
- Iteration # 1 Level # 3 Size 32x 32x 1
- Iteration # 2 Level # 3 Size 32x 32x 1
- Iteration # 3 Level # 3 Size 32x 32x 1
- Iteration # 4 Level # 3 Size 32x 32x 1
- Iteration # 5 Level # 3 Size 32x 32x 1
- Iteration # 6 Level # 3 Size 32x 32x 1
- Iteration # 7 Level # 3 Size 32x 32x 1
- Iteration # 8 Level # 3 Size 32x 32x 1
- Iteration # 9 Level # 3 Size 32x 32x 1
- Iteration # 10 Level # 3 Size 32x 32x 1
- processing level # 2
- Iteration # 1 Level # 2 Size 64x 64x 1
- Iteration # 2 Level # 2 Size 64x 64x 1
- Iteration # 3 Level # 2 Size 64x 64x 1
- Iteration # 4 Level # 2 Size 64x 64x 1
- Iteration # 5 Level # 2 Size 64x 64x 1
- Iteration # 6 Level # 2 Size 64x 64x 1
- Iteration # 7 Level # 2 Size 64x 64x 1
- Iteration # 8 Level # 2 Size 64x 64x 1
- Iteration # 9 Level # 2 Size 64x 64x 1
- Iteration # 10 Level # 2 Size 64x 64x 1
- processing level # 1
- Iteration # 1 Level # 1 Size 128x 128x 1
- Iteration # 2 Level # 1 Size 128x 128x 1
- Iteration # 3 Level # 1 Size 128x 128x 1
- Iteration # 4 Level # 1 Size 128x 128x 1
- Iteration # 5 Level # 1 Size 128x 128x 1
- Iteration # 6 Level # 1 Size 128x 128x 1
- Iteration # 7 Level # 1 Size 128x 128x 1
- Iteration # 8 Level # 1 Size 128x 128x 1
- Iteration # 9 Level # 1 Size 128x 128x 1
- Iteration # 10 Level # 1 Size 128x 128x 1
Compose and apply transformations to floating images¶
We now have the initial rigid, affine & non-linear transformations to register each floating image onto the reference image. We thus need to compose them (as we used the left_trsf parameter) before applying them to their respective floating image.
res_imgs = [ref_img] # initialize images list (to average)
for i, flo_img in enumerate(img_list[1:]):
# Compose the linear and non-linear transformations (initial + affine + vectorfield)
comp_trsf = compose_trsf([init_trsfs[i], lin_trsfs[i], vf_trsfs[i]], template_img=ref_img)
# Apply the composed transformation to the floating image
res_imgs.append(apply_trsf(flo_img, comp_trsf, interpolation='linear'))
print("Registered image shapes:")
print("\n".join([f"Image #{n}: {im.shape}" for n, im in enumerate(res_imgs)]))
Registered image shapes:
Image #0: (184, 184)
Image #1: (184, 184)
Image #2: (184, 184)
We can see that all images now have the reference image shape.
print("Registered image voxelsizes:")
print("\n".join([f"Image #{n}: {im.voxelsize}" for n, im in enumerate(res_imgs)]))
Registered image voxelsizes:
Image #0: [0.5, 0.5]
Image #1: [0.5, 0.5]
Image #2: [0.5, 0.5]
WARNING: We should see that all images now have the reference image voxelsize.
Visualisation of the final registration¶
Then, we represent one slice of the intensity signal of each image in a different color to appreciate the “signal alignment” (or not).
The corresponding color code:
reference
090223-p58-flo-top.lsm: redregistered
090223-p58-flo-tilt1.lsm: greenregistered
090223-p58-flo-tilt2.lsm: blue
z_slices = [5, 10, 15, 20]
fig, axes = plt.subplots(1, 4)
fig.set_size_inches(15, 5)
fig.set_dpi(160)
fig.suptitle("Final multi-angle registration")
blend = combine_channels(res_imgs, colors=['red', 'green', 'blue'])
for n, zsl in enumerate(z_slices):
imshow(axes[n], blend[zsl, :, :], title = f"z-slice #{zsl}")
fig.subplots_adjust(wspace=0, hspace=0)
fig.tight_layout()
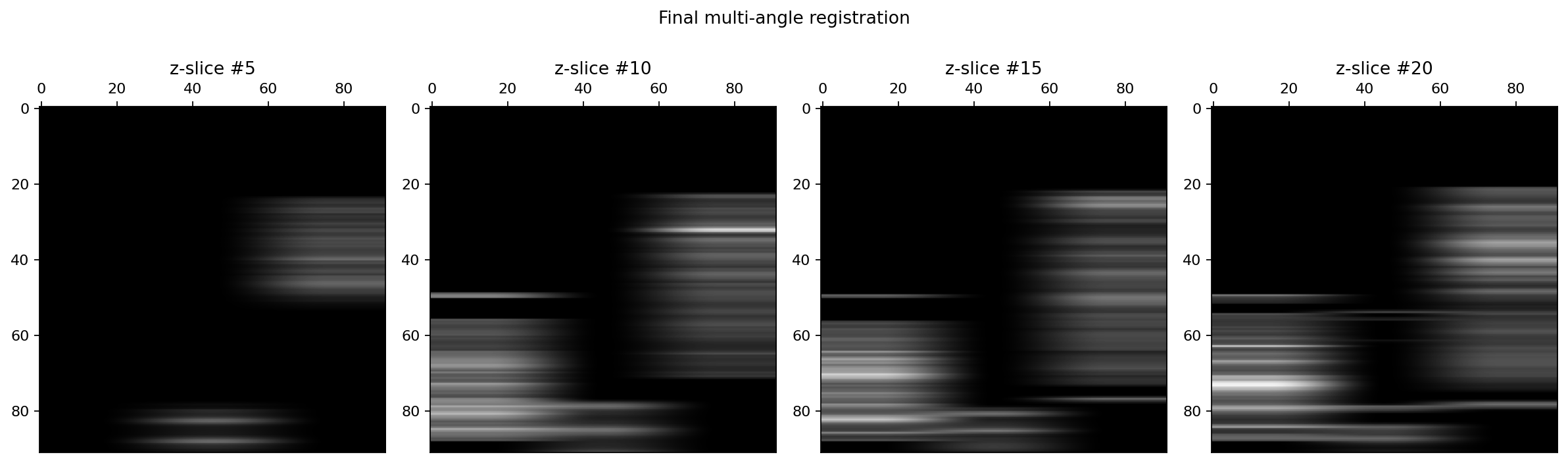
Combine the reference and transformed images with specified fusion method:¶
We start by computing the mean image from the reference image and the set of floating image registered on the reference.
mean_image = images_averaging(res_imgs, method='mean')
print("Mean registered image shape:")
print(mean_image.shape)
Mean registered image shape:
(184, 184)
It’s the same shape as the reference image since it was used as template!
print("Mean registered image voxelsize:")
print(mean_image.voxelsize)
Mean registered image voxelsize:
[0.5, 0.5]
It’s the same voxelsize as the reference image since it was used as template!
Let’s have a look at a few slices…
z_slices = [5, 10, 15, 20]
fig, axes = plt.subplots(1, 4)
fig.set_size_inches(15, 5)
fig.set_dpi(160)
fig.suptitle("Mean image")
for n, zsl in enumerate(z_slices):
imshow(axes[n], mean_image.get_slice(zsl), title = f"z-slice #{zsl}")
fig.subplots_adjust(wspace=0, hspace=0)
fig.tight_layout()
---------------------------------------------------------------------------
KeyError Traceback (most recent call last)
Cell In[28], line 8
6 fig.suptitle("Mean image")
7 for n, zsl in enumerate(z_slices):
----> 8 imshow(axes[n], mean_image.get_slice(zsl), title = f"z-slice #{zsl}")
10 fig.subplots_adjust(wspace=0, hspace=0)
11 fig.tight_layout()
File /builds/J-gEBwyb/0/mosaic/timagetk/src/timagetk/components/spatial_image.py:1355, in SpatialImage.get_slice(self, slice_id, axis)
1301 def get_slice(self, slice_id, axis='z'):
1302 """Return a SpatialImage with only one slice for given axis.
1303
1304 Parameters
(...)
1353
1354 """
-> 1355 arr, axes_order, ori, vxs, md = self._get_slice(slice_id, axis)
1356 return SpatialImage(arr, origin=ori, voxelsize=vxs, metadata=md, unit=self.unit, axes_order=axes_order)
File /builds/J-gEBwyb/0/mosaic/timagetk/src/timagetk/components/spatial_image.py:1241, in SpatialImage._get_slice(self, slice_id, axis)
1239 # -- Correspondence between `axis` string and `axis_idx` index:
1240 if isinstance(axis, str):
-> 1241 axis_idx = self.axes_order_dict[axis.upper()]
1242 else:
1243 axis_idx = cp.copy(axis)
KeyError: 'Z'
Centering the mean image in the reference frame:¶
To avoid bias toward the chosen reference image and “center” the final fused image between the geometry of the three images, we invert the composed affine and non-linear deformation.
trsfs2average = []
trsfs2average.append(create_trsf('identity', template_img=ref_img, trsf_type='vectorfield'))
for i, flo_img in enumerate(img_list[1:]):
trsfs2average.append(compose_trsf([lin_trsfs[i], vf_trsfs[i]], template_img=ref_img))
mean_vf_trsf = trsfs_averaging(trsfs2average, trsf_type="vectorfield")
from timagetk.algorithms.template import iso_template
super_rez = True
if super_rez:
# Resample the reference image in the template image:
template = iso_template(ref_img, method="min")
else:
template = ref_img
Apply the invert of the combined non-linear deformations to the combined image¶
inv_vf_trsf = inv_trsf(mean_vf_trsf, template=ref_img)
BAL_Inverse3DVectorField: divergence: 0/1997504, non-convergence: 65/1997504
Note:
using the
mean_imageastemplate_imgininv_trsfled to a weird fused image with inverted shape!do NOT forget to add the
template=ref_imgwith a vectorfield transformation
fused_img = apply_trsf(mean_image, inv_vf_trsf, interpolation='linear')
Let’s have a look at a few slices…
cuts = [1 / 5., 1 / 3., 1 / 2., 2 / 3.]
z_sh = fused_img.get_shape('z')
z_slices = [int(round(c * z_sh, 0)) for c in cuts]
fig, axes = plt.subplots(1, 4)
fig.set_size_inches(15, 5)
fig.set_dpi(160)
fig.suptitle("Fused image")
for n, zsl in enumerate(z_slices):
imshow(axes[n], fused_img.get_slice(zsl), title = f"z-slice {zsl}/{z_sh}")
fig.subplots_adjust(wspace=0, hspace=0)
fig.tight_layout()
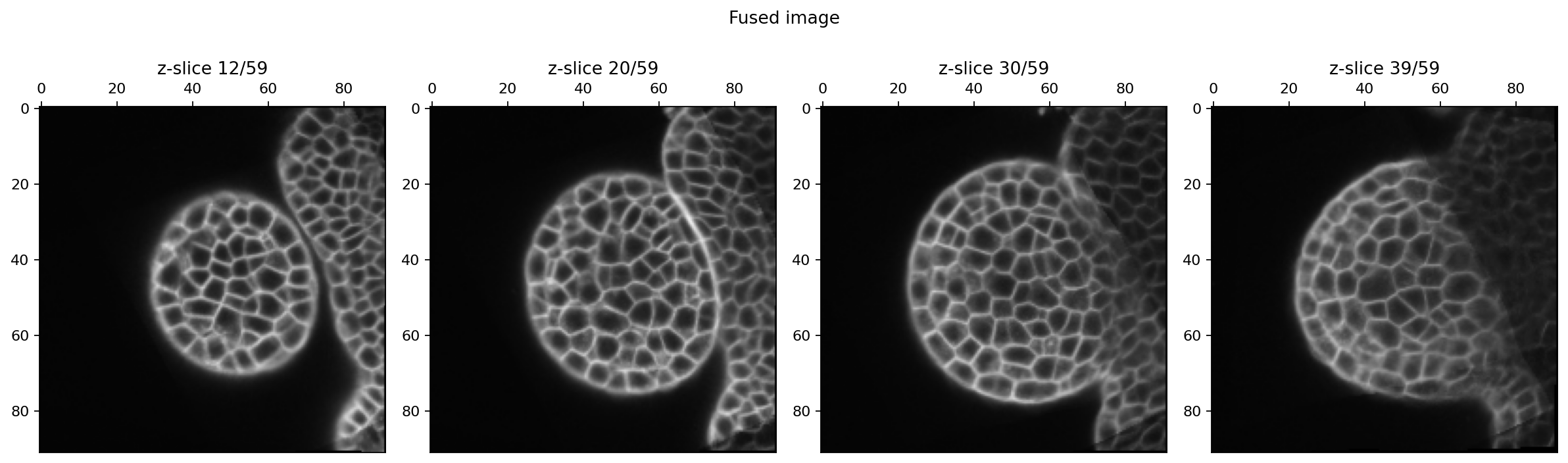

We have succesfully obtained a fused image from the chosen reference image!
It is now possible to iterate this procedure using the obtained fused image at iteration i as reference for the iteration i+1!
